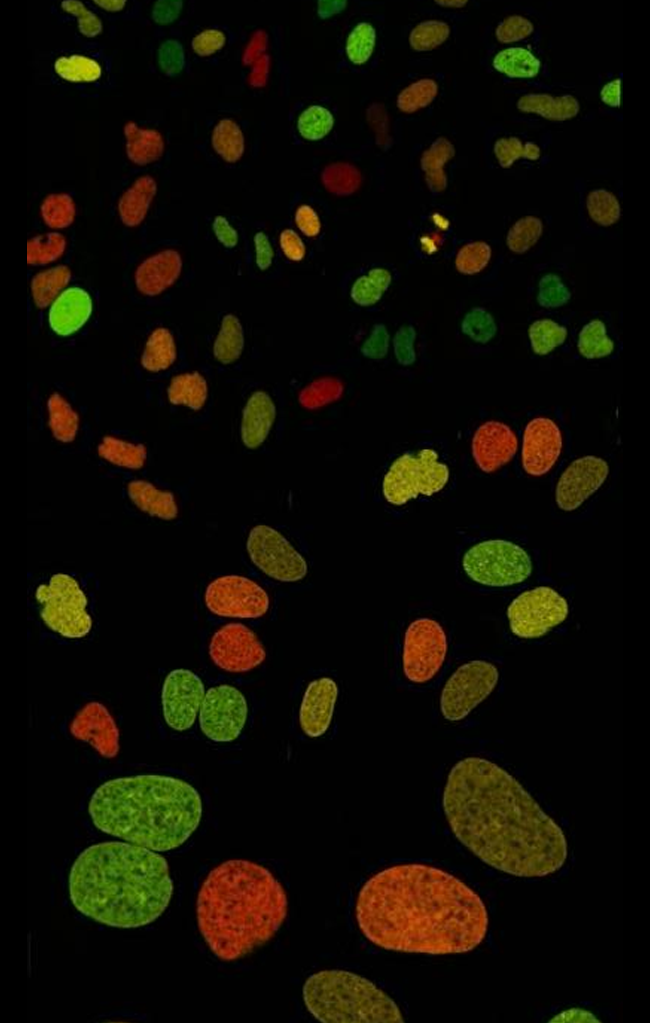
Liz has developed a method—microscopy combined with fluorescence fluctuation analysis—to track the movement of molecules around the complex DNA networks within the nuclei of live cells.
Inside the nucleus, DNA repair machines find sites of damage to prevent genetic mutations, and transcription factors search for target DNA sequences to maintain normal gene expression.
At any given time, thousands of molecules move around the nucleus. Building a complete road map for navigating the DNA network will help to understand how and why deregulated nuclear traffic leads to diseases like cancer.
By analysing changes in fluorescence intensities, Liz discovered that DNA networks rearrange to create highways for repair and transcription factors to arrive at target destinations.
About Elizabeth Hinde
In 2010 Elizabeth completed her PhD in fluorescence spectroscopy at the University of Melbourne and was then recruited to the University of California, Irvine, USA to pursue a post-doctoral fellowship under the mentorship of Professor Enrico Gratton. In the Gratton lab (2010-2013) Elizabeth developed methods based on fluorescence correlation spectroscopy (FCS) and fluorescence lifetime imaging microscopy (FLIM) to quantify chromatin dynamics in live cells. With the aim of applying this technology to cell biology, Elizabeth returned to Australia in 2013 as a UNSW Vice Chancellor Fellow under the mentorship of Professor Katharina Gaus. At UNSW (2013-2015) Elizabeth’s research revealed insights into genome organisation that led to the award of a Cancer Institute of NSW Early Career Research Fellowship in 2015 and this enabled her to establish a research group within the EMBL Australia node for Single Molecule Science. Liz has relocated to Bio21, but maintains close collaborations with SMS.
Liz is now at Bio21 Institute